FHIR Server powers Project Vulcan @ HL7 FHIR Connectathon

Put the InterSystems FHIR Server to work at the HL7 FHIR Connectathon in Baltimore last weekend, posting bundles, getting all RESTy with resources, and interrogating the Vulcan Implementation Guide. We straddled the Real World Data (RWD) and Schedule of Activity (SoA) projects for Vulcan IG folks moving the ball forward connecting clinical research and healthcare data. We employed a pretty decent approach to meet the requirements,
Myself along with InterSystems peeps (Huy, Russell, Regilo), were on the hook to support the FHIR Server and data appropriation for the event, if you have ever been tasked with typing up FHIR Bundles for a specific use case, I think you can appreciate how elusive the difficulty can be.
We provisioned a couple of ghetto blaster sized InterSystems FHIR Servers, 16 core/128GB, one for each project and went to work for each team. Teams came prepared with their Python Notebooks against our endpoints, with the expectation of writing a FHIR Story for ACS with a procedural test script validating the calls returned with the anticipated results.
The cohort target for RWD read quite simple in the example queries for Acute Coronary Syndrome to me:
Right?
/Patient?birthdate=le2002-09-01&gender=male,female
/Encounter?reason-code:below=I20,I21,I22,I23,I24,I25&date=ge2020-09-01&date=le2021-09-31&status=finished&dischargeDisposition:not=exp
/MedicationAdministration?status=completed&effective-time=ge[Encounter-Start-Date]&
code=http://www.nlm.nih.gov/research/umls/rxnorm|1116632,http://www.nlm.nih.gov/research/umls/rxnorm|613391,http://www.nlm.nih.gov/research/umls/rxnorm|32968,http://www.nlm.nih.gov/research/umls/rxnorm|687667,http://www.nlm.nih.gov/research/umls/rxnorm|153658
Sure...
Fails Synthea SubModule Bundle Bomb Though hopeful, our first hail mary was to generate 2 million resources and 1000's of Synthea Bundles (using this sweet InterSystems repo to do it) to the transaction endpoint and the Data Management upload facility including a bunch of heart centric submodules, and fancy switches...
docker run --rm -v $PWD/output:/output -v $PWD/modules:/modules --name synthea-docker intersystemsdc/irisdemo-base-synthea:version-$VERSION --exporter.practitioner.fhir.export true --exporter.hospital.fhir.export true --exporter.fhir.use_us_core_ig true -p 500 -s 21 -d /modules
This fell short, not surprising I guess, but I kind of hoped we would at least get close. We did not.
Wins The Data Generation Process So Geoff Low clued us in on his process, and we forked and were off and running.... this is how the process goes.
-
Grab Yourself a starting point bundle, particularly one that is relevant to Research and that has patients and encounters at a minimum, uses the SDTM bundle out here on sourceforge:
-
Now, implement a patching step, to fix the bundle in place, ensure it loads successfully in the FHIR Server, upload or bundle POST.
-
Once we have a prestine starting point, augment the encounters with the anticipated encounter reason codes ( I20,I21,I22,I23,I24,I25 ).
reasons = ["I21","I22","I23","I24","I25"]```
- Add medications to the encounters, include all the expected medications for the encounter for the ACS Research Study.
medchoices = [
{
"code": "1116632",
"display": "ticagrelor"
},
{
"code": "613391",
"display": "prasurgrel"
},
{
"code": "32968",
"display": "clopidogrel"
}
]
medchoice = random.choice(medchoices)
medconcept = CodeableConcept(
coding=[
Coding(
code=medchoice["code"],
display=medchoice["display"],
system="http://www.nlm.nih.gov/research/umls/rxnorm",
)
]
)
IMPORTANT! Make sure the medications fall within the dates/times of the encounter.
- Next, make sure the Encounters are indeed hospitalizations, and include all the applicable types.
statuses = ["in-progress","finished"]
dischargeCodes =
{
"code": "home",
"display": "Home"
},
{
"code": "hosp",
"display": "Hospice"
},
{
"code": "exp",
"display": "Expired"
},
{
"code": "long",
"display": "Long-term care"
},
{
"code": "alt-home",
"display": "Alternative home"
},
]
Another Win Implementing the IG Search Parameter The Vulcan Implementation Guide includes a search parameter that is required to return encounters by discharge disposition, we approached this in two ways:
a. Load the entire IG b. Load the Search Parameter
Loading the Search Parameter by itself was our shortest path and we'd been in the hotseat long enough, so here is how that process went.
Created a folder /tmp/mypackage, add the search parameter from the IG, and your own package file.
[irisdeploy@ip-192-168-0-37 tmp]$ tree /tmp/mypackage
/tmp/mypackage
├── package.json
└── parameter.json
Your package file can look a little something like this:
{
"name":"ron.sweeney.r4",
"version":"0.0.1",
"dependencies": {
"hl7.fhir.r4.core":"4.0.1"
}
}
Next load that up into IRIS like so:
TL4:IRIS:FHIRDB>do ##class(HS.FHIRMeta.Load.NpmLoader).importPackages($lb("/tmp/mypackage"))
Saving ron.sweeney.r4@0.0.1
Load Resources: ron.sweeney.r4@0.0.1
Next step is to associate the package to the endpoint:
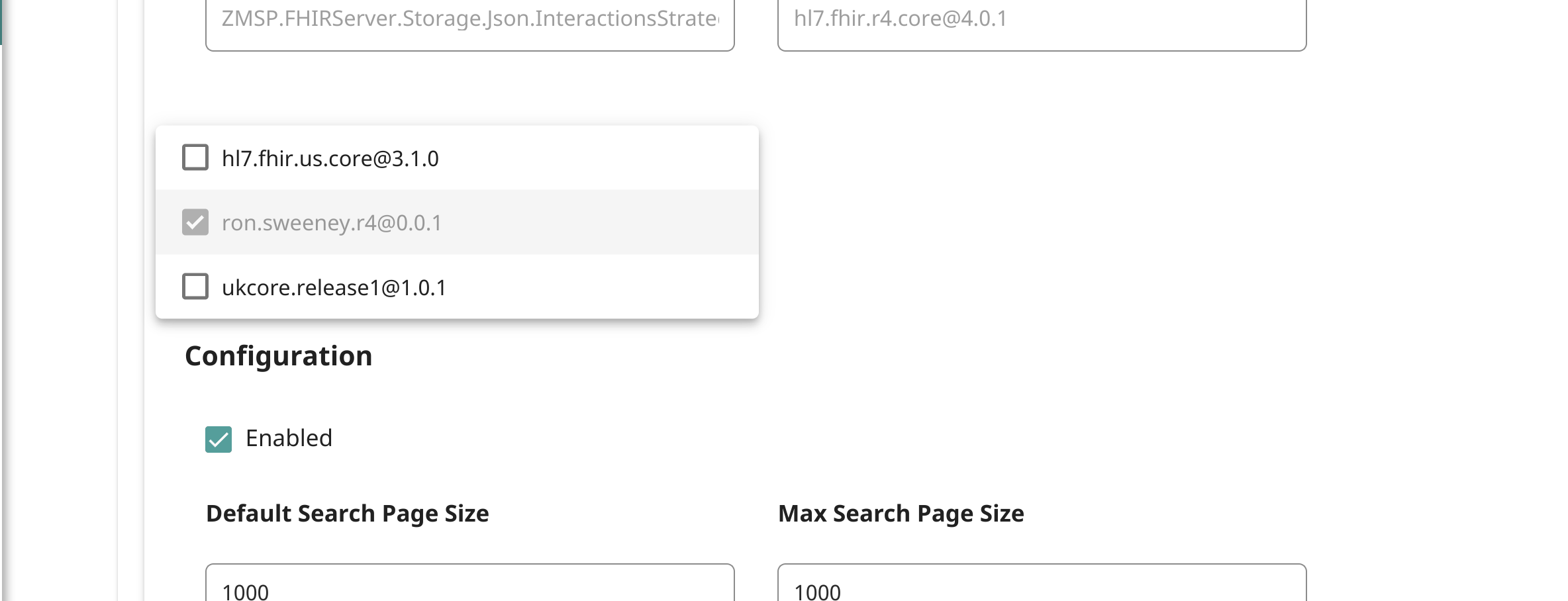
Then, re-index the encounters in the repository:
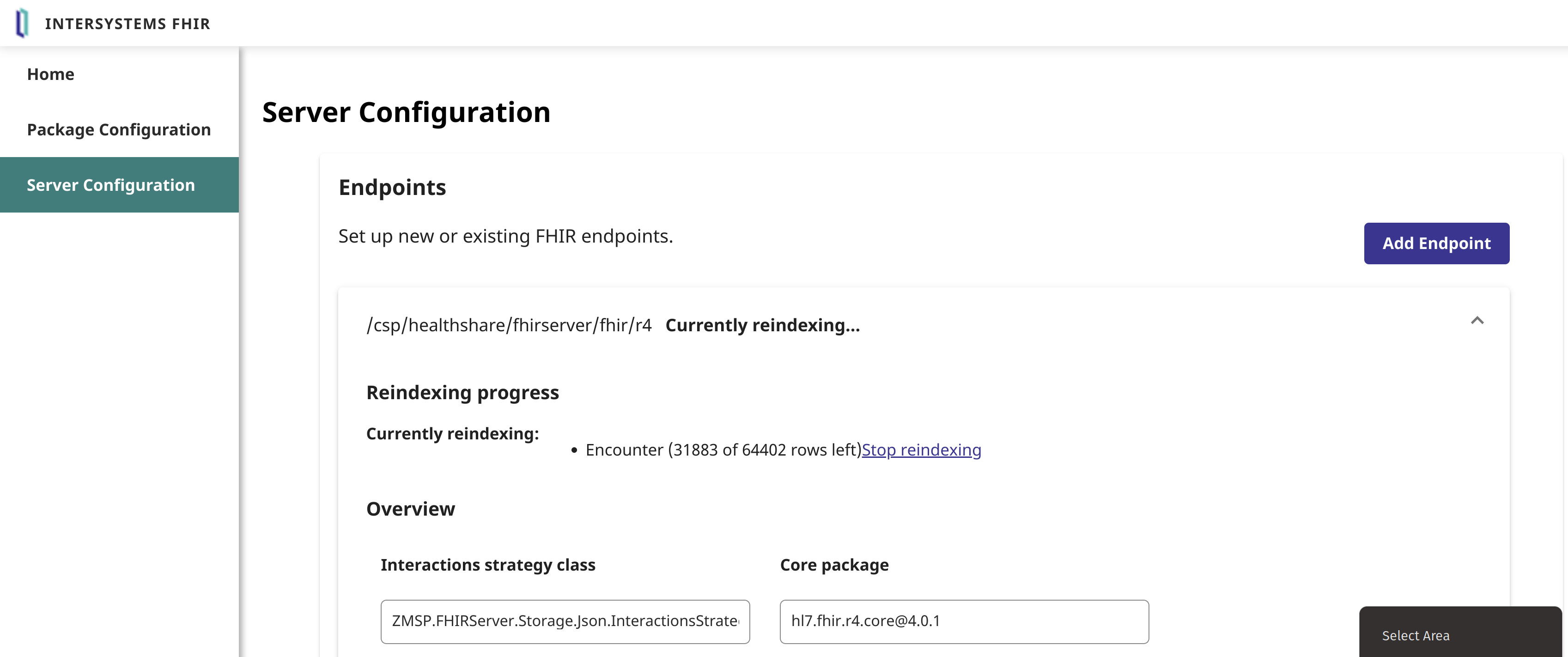
You should now have the search parameter/package loaded on your endpoint:
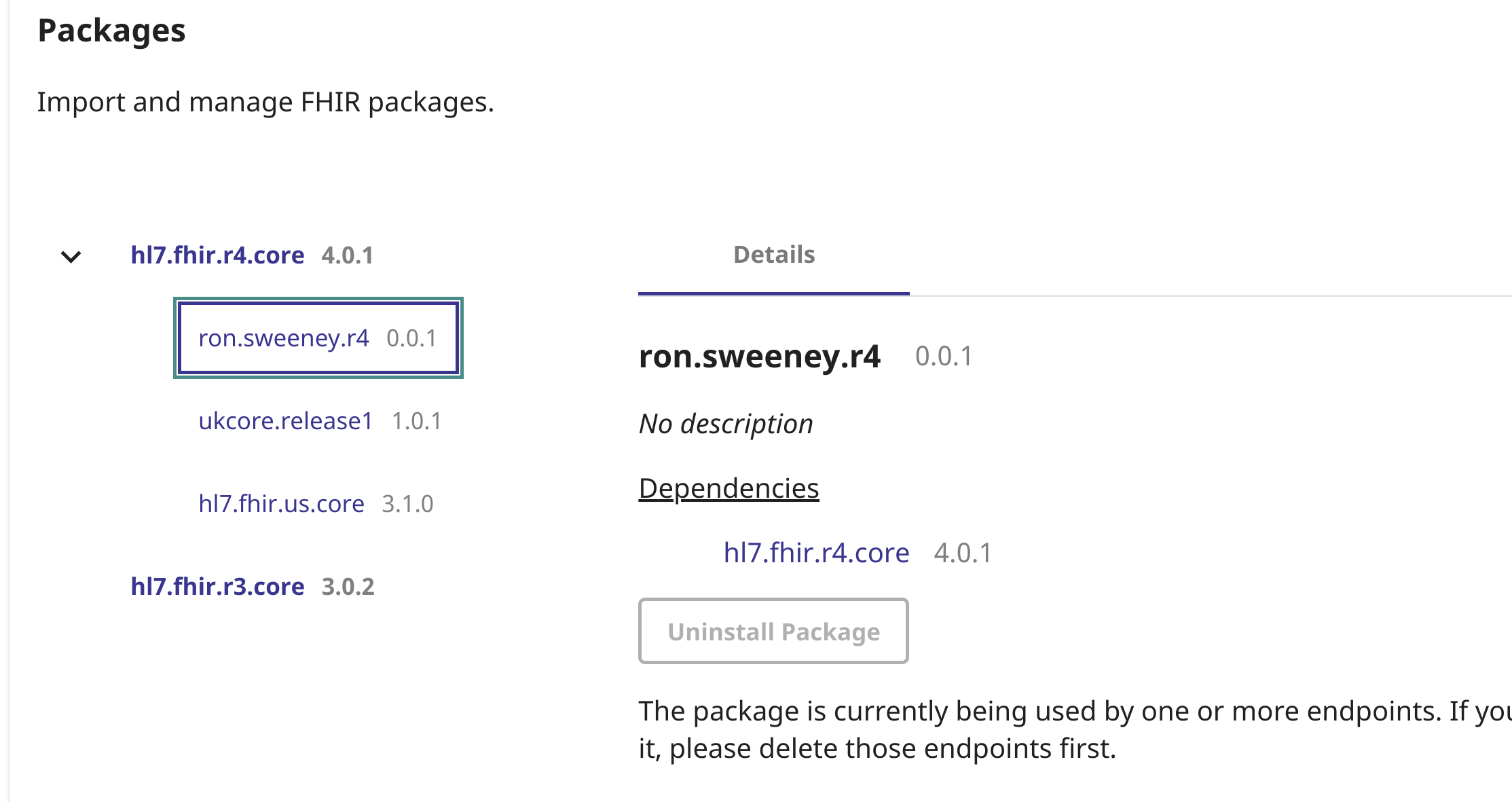
With the search parameter loaded, we were now able to utilize it like so!
curl https://fhir.ggyxlz8lbozu.workload-prod-fhiraas.isccloud.io/Encounter?dischargeDisposition=hosp
Anyway, the process worked for us, and approaching bundle generation through the eyes of Python was a way better approach than Synthea sometimes for exacting datasets.
The resulting dataset we walked away with Sunday is here.
Just wanted to share the experience and thank the teams from the projects for making what I thought would be a nightmare experience fun and educational!
Comments
this is fantastic! thank you so much for supporting the event. building out appropriate data set to test IG is a critical step.
What a great feedback, thanks for sharing this with us.